Oteng’o, Won, and Cho: Fissipedicella orientalis gen. et sp. nov. (Ralfsiales, Phaeophyceae), a new crustose brown alga from Korea based on molecular and morphological analyses
ABSTRACT
The crustose brown algal family Ralfsiaceae comprises four genera: Analipus, Endoplura, Heteroralfsia, and Ralfsia. This study provides a detailed description of Fissipedicella orientalis gen. et sp. nov. based on molecular and morphological analyses. Our phylogenetic analyses from rbcL and concatenated dataset (rbcL + 5′ region of cytochrome c oxidase subunit I [COI-5P]) reveal that specimens collected in Korea are nested in a distinct new clade within Ralfsiaceae with robust bootstrap support and Bayesian posterior probabilities. The sequence divergences for rbcL and COI-5P between F. orientalis and other genera within Ralfsiaceae are 7.4–10.1 and 17.5–21.2%, respectively. Fissipedicella orientalis is characterized by crustose thalli with a hypothallial basal layer and erect perithallial filaments, tufts of hairs in pits, a single chloroplast per cell, plurangia with 1–3 sterile cells, and unangia on stalks composed of 1–6 vertically or obliquely cleaved cells. We propose that F. orientalis can be recognized as a new genus-level taxon within Ralfsiaceae, even though a single species represents it. Our new genus, Fissipedicella, is distinguished from the other members within the Ralfsiaceae by the type of thallus, the number of chloroplasts and tufts of hairs in pits, and the development of unangia.
Key words: COI-5P; Fissipedicella; phylogeny; Ralfsiaceae; rbcL, taxonomy
INTRODUCTION
The family Ralfsiaceae Farlow was first described by Farlow (1881). It was characterized by: (1) rounded to irregularly spreading crust developed from a discoid germination stage; (2) a basal layer of radiating appressed filaments producing simple or slightly branched and tightly appressed erect or assurgent vegetative filaments; (3) single or few chloroplasts without a pyrenoid per cell; (4) intercalary plurangial reproductive structures terminated by sterile cell(s); (5) unangia with paraphysis on the terminal part of erect filaments; (6) diplohaplontic and iso- or heteromorphic life history ( Farlow 1881, Setchell and Gardner 1924, Abbott and Hollenberg 1976, Womersley 1987, Parente and Saunders 2019). Currently, Ralfsiaceae is composed of four genera: Analipus Kjellman, Endoplura Hollenberg, Heteroralfsia Kawai, and Ralfsia Berkeley ( Lim et al. 2007, Silberfeld et al. 2014, Parente and Saunders 2019, Guiry and Guiry 2023). Although the principal morphological characters used to distinguish genera within the Ralfsiaceae are known to be the type of thallus, the number of plastids per cell, the number of sterile cells on plurangia, and the development of unangia ( Tanaka and Chihara 1981b, Lim et al. 2007, León-Alvarez et al. 2017, Oteng’o et al. 2021, Parente et al. 2021), it is challenging to accurately identify these crustose species based on traditional morpho-anatomical characteristics because they lack robust taxonomic features, particularly in sterile samples ( Parente and Saunders 2019). Recently, molecular analyses using rbcL have been utilized to infer the phylogenetic relationships among genera within Ralfsiaceae ( Lim et al. 2007, Silberfeld et al. 2014, Parente and Saunders 2019, Oteng’o and Won 2020). Most crustose brown algae in Korea have traditionally been identified mainly as Ralfsia or Neoralfsia species based on their morphology ( Lee and Kang 1986, 2001, Lee 2008, Keum 2010). However, recent research by Oteng’o et al. (2021, 2022) introduced two new Endoplura species, E. jejuensis A. O. Oteng’o, T. O. Cho & B. Y. Won and E. koreana A. O. Oteng’o, T. O. Cho & B. Y. Won and a new genus called Sungminia Oteng’o, Won & T. O. Cho within Sungminiaceae Oteng’o, Won & T. O. Cho from Korea based on morphological and molecular analyses. These findings showed the possibility that Korea may be one of the hotspots for crustose brown algal biodiversity. We collected several unidentified samples of Ralfsia-like crusts from intertidal areas along Korean coastal shores. This study used morphological characterizations and examined their phylogenetic relationships through rbcL and 5′ region of cytochrome c oxidase subunit I (COI-5P) gene sequences. As a result, we propose Fissipedicella orientalis gen. et sp. nov., which is assigned to the family Ralfsiaceae.
MATERIALS AND METHODS
DNA extraction and amplification
From 2018 to 2020, we collected samples from intertidal areas along the coastlines of Korea ( Supplementary Table S1). Genomic DNA was extracted using the NucleoSpin Plant II Kit from Macherey-Nagel (Düren, Germany), following the manufacturer’s instructions. The rbcL and COI-5P were amplified using polymerase chain reaction with HelixAmp Ready-2x-Go premix from NanoHelix Co., Ltd. (Daejeon, Korea), following the manufacturer’s protocol. The primer sets used for the amplifying rbcL and COI-5P were as specified in a previous study by Oteng’o et al. (2021).
Sequence analyses
The sequence datasets were assembled by combining previously published sequences from GenBank with 12 newly generated rbcL and 11 COI-5P sequences from this study ( Supplementary Table S1). We analyzed two datasets: one consisting of 57 rbcL sequences and the other with 57 concatenated sequences ( rbcL + COI-5P). We considered sequences that were unavailable in GenBank as missing data. We included sequences from Cladostephus spongiosus (Hudson) C. Agardh and Microzonia phinneyi (E. C. Henry & D. G. Müller) Camacho & Fredericq for the outgroup. The sequences were aligned using ClustalW ( Thompson et al. 1994) and manually edited using Geneious Prime (v.2023.0.1; Biomatters Ltd., Auckland, New Zeland). To determine the best partition scheme and model evolution, we used PartitionFinder 2 ( Lanfear et al. 2016). For the maximum likelihood (ML) analysis, RAxMLGUI v1.5 ( Silvestro and Michalak 2012) was employed with the General Time Reversible (GTR) + Γ + I model with 1,000 bootstrap replications. We conducted Bayesian inference using MrBayes 3.2.6 ( Huelsenbeck and Ronquist 2001, Ronquist and Huelsenbeck 2003). Markov chain Monte Carlo runs were carried out for 2,000,000 generations, with one cold chain and three heated chains, using the GTR + Γ + I evolutionary model. Trees were sampled every 1,000 generations. Summary trees were generated with a burn-in value of 25%. The convergence was assessed using Tracer v.1.7.2 ( Rambaut et al. 2018). Phylogenetic trees were constructed using the ML method in RAxML and MrBayes and visualized FigTree v.1.4.0 ( Rambaut 2012). Genetic distances were calculated using the p-distance method in Mega X ( Kumar et al. 2018).
Morphological analyses
To examine external morphological characteristics, samples were photographed using a waterproof digital camera (Nikon COOLPIX AW100; Nikon Corp., Tokyo, Japan). Before morphological observation, the samples were carefully detached from the substrate using a single-edged blade. We either squashed or prepared microtome sections of portions of each sample mounted on microscopy slides to observe internal morphology. The collected samples were embedded in a matrix (OCT; CellPath Ltd., Newtown, Wales, UK) to prepare microtome-sectioned samples. These embedded samples were then sectioned to a thickness of 8–10 μm using a freezing microtome (Shandon Cryotome FSE; Thermo Shandon Ltd., Loughborough, UK). The sliced and squashed samples underwent staining using a 1 : 1 combination of aqueous aniline blue and acetic acid for better visualization and analysis. The staining process enhanced the visibility of certain structures and details. The stained sections were mounted in 50% corn syrup and observed under a microscope. Photographs of samples were taken using a DP-71 camera (Olympus, Tokyo, Japan) mounted on a BX-51TRF microscope (Olympus). To ensure clarity and improve visual presentation, the digitized images were edited using Adobe Photoshop software v.6.1 (Adobe Systems Inc., San Jose, CA, USA). Representative voucher specimens examined in this study were deposited in the herbarium of Chosun University (CUK) and the National Institute of Biological Resources (NIBR), Korea.
RESULTS
Phylogenetic analyses
A total of 12 rbcL sequences (984 bp) and 11 COI-5P (654 bp) sequences were generated from newly collected samples from Korea ( Supplementary Table S1). The phylogenetic tree inferred from concatenated dataset ( rbcL + COI-5P) indicated that the unidentified Ralfsia-like crustose algae from Korea form a distinct and previously unknown genus, Fissipedicella orientalis gen. et sp. nov., within the family Ralfsiaceae ( Fig. 1). Phylogenetic analyses based on the rbcL sequences showed overall agreement with concatenated phylogeny ( Supplementary Fig. S1). Through our multi-gene sequence analyses, significant genetic differences were observed in rbcL (7.4–8.9%) and COI-5P (17.5–19.3%) sequences between Fissipedicella and Ralfsia within Ralfsiaceae ( Table 1).
Morphological observations
Fissipedicella A. O. Oteng’o, B. Y. Won & T. O. Cho gen. nov
Description
Thalli are crustose with circular to irregular outlines, firmly attached to the substratum, and lacking rhizoids. They comprise two main parts: the hypothallial basal layer and the perithallial erect filament. The hypothallial basal layer is formed by one to several prostrate filaments, which are made up of elongated cells that give rise to perithallial erect filaments. The perithallial erect filaments exhibit branching, straight, or upward curvature, gradually tapering towards the surface, and are firmly adjoined. The chloroplast is parietal and one per cell. Tufts of hair in pits frequently arise from the lower to middle portions of the perithallial erect filaments. The plurangial reproductive structure is intercalary, composed of 1–2 reproductive filaments, and terminated by 1–3 sterile cells. Unangia originate from perithallial erect cells that branch to produce the paraphysis and stalk. The stalk serves as a supportive structure for the terminal unangium. The stalks are composed of vertically or obliquely cleaved cells.
Type species
Fissipedicella orientalis.
Etymology
The name “Fissipedicella” refers to unangial pedicels (stalks) formed by vertical cell division.
Korean name
갯바위딱지속.
Fissipedicella orientalis A. O. Oteng’o, B. Y. Won & T. O. Cho sp. nov. (Fig. 2)
Description
Thalli are epilithic crusts, exhibiting a range of colors from greenish or reddish-brown to dark brown, which become very dark brown when dry. They have a circular to irregular outline and are often mere together, with no distinct lighter margins ( Fig. 2A & B). Thalli are 0.2–4.5 (−7.5) cm across, 85–421 μm thick, firmly attached to the substratum, with inconspicuous growth lines and mostly smooth surface, with rust-red undersurface, and lacking rhizoids ( Fig. 2A & B). The hypothallial basal layer comprises one to several cell layers in which cells are 4–12 μm wide and have a width-to-length ratio of 1 : 1.8–3.8, giving rise to erect perithallial filaments ( Fig. 2C & D). The erect perithallial filaments are branched, straight, or upwardly curved, tapering towards the surface, and firmly adjoined with one another to form pseudoparenchymatous tissue ( Fig. 2D). Cells of the erect perithallial filaments are 4–25 μm long and 4–16 μm wide. The chloroplast is parietal and one per cell ( Fig. 2E). Tufts of hair in pits arise from the basal to mid parts of erect perithallial filaments ( Fig. 2F). Reproductive portions of the plurangial and unangial sori form slightly elevated areas on the same or different thalli ( Fig. 2G & J). Plurangial reproductive structures are intercalary, 43–202 μm long, composed of 1–2 reproductive filaments, and terminated by 1–3 sterile cells ( Fig. 2H & I). Unangial reproductive thalli give rise to produce two types of filaments, both originating from erect perithallial cells. These filaments branch out to form paraphyses, and there are also shorter biseriate stalk cells that support terminal unangia ( Fig. 2K & L). Unangia are obovoid to ellipsoid to oblong, 47–116 μm long, and 7–40 μm wide, and terminally produced on 1–6 celled stalks that are vertically or obliquely cleaved ( Fig. 2L). The paraphyses are clavate, 92–148 μm long, 2–12 μm wide, and composed of 5–10 cells ( Fig. 2L).
Holotype
CUK18754, Mar 3, 2018, coll. by B. Y. Won and T. O. Cho deposited in the CUK herbarium, Korea.
Type locality
Pado-ri, Sowon-myeon, Taean-gun, Chungcheongnam-do, Korea (36°43′05.65″ N, 126°07′ 33.79″ E).
Isotypes
NIBROR0000004864 (CUK18765) at the National Institute of Biological Resources (NIBR), Korea.
Etymology
The specific epithet, orientalis, refers to the Orient (Far East Asia) region where the alga grows naturally.
DNA sequences of type specimens
For the holotype, OK031043 (rbcL) and OK031055 (COI-5P).
Paratypes
CUK18821, CUK18822, CUK18825 (Apr 1, 2018, Jeongdo-ri, Wando-eup, Wando-gun, Jeollanam-do, Korea); CUK19178A (Oct 8, 2018, Gijang, Ilgwang-myeon, Gijang-gun, Busan, Korea); CUK19219B (Nov 3, 2018, Pado-ri, Sowon-myeon, Taean-gun, Chungcheongnam-do, Korea); CUK19509C, CUK19510C, CUK19512B, CUK19519 (Apr 19, 2019, Seongsan Ilchul Peak, Seongsan-eup, Seogwipo-si, Jeju Island, Korea); CUK19606C (May 1, 2019, Seongsan Ilchul Peak, Seongsan-eup, Seogwipo-si, Jeju Island, Korea); CUK19696, CUK19699, CUK19703, CUK19706, CUK19707B, CUK19709, CUK19710 (Jun 6, 2019, Yeraehaean-ro, Seogwipo-si, Jeju Island, Korea); CUK20040D, CUK20041D (Oct 26, 2019, Bomok Port, Bomok-dong, Seogwipo-si, Jeju Island, Korea); CUK20066A (Oct 28, 2019, Hwasam-ri, Yongnam-myeon, Tongyeong-si, Gyeongsangnam-do, Korea); CUK20331, CUK20332 (May 8, 2020, Jeongdo-ri, Wando-eup, Wando-gun, Jeollanam-do, Korea); CUK20753A, CUK20754, CUK20755, CUK20759E, CUK20760A (Aug 17, 2020, Seongsan Ilchul Peak, Seongsan-eup, Seogwipo-si, Jeju Island, Korea); CUK20771, CUK20772 (Oct 17, 2020, Dolsan Port, Dolsan-eup, Yeosu-si, Jeollanam-do, Korea).
Reproductive phenology
Plants with both plurangial and unangial reproductive structures have been collected during March to June and also in November.
Distribution and habitat
This species has been observed in various localities along the western coast (Taean), southern coast (Busan, Tongyeong, and Wando), and Jeju Island, Korea. It is found on hard substrates, including pebbles and rocks in the intertidal zone.
DISCUSSION
Based on morphological and molecular analyses, we propose a new genus and species, Fissipedicella orientalis gen. et sp. nov. The sequences of both rbcL and concatenated dataset ( rbcL + COI-5P) from specimens collected in Korea were nested in a distinct clade within Ralfsiaceae and were distinguished from the other members within the Ralfsiaceae. Fissipedicella orientalis has a sister relationship with Endoplura and an unidentified clade (incertae sedis) ( Fig. 1, Supplementary Fig. S1). Although Endoplura shows a sister relationship with our F. orientalis in the phylogeny based on both rbcL and concatenated dataset ( rbcL + COI-5P), it is distinguished from F. orientalis by having several chloroplasts per cell and sessile unangia ( Hollenberg 1969, Tanaka and Chihara 1981b, Lim et al. 2007, Oteng’o et al. 2021). Ralfsia within the Ralfsiaceae also distinguished from F. orientalis by having unangia either sessile or on stalks of 1–2 horizontally cleaved cells and plurangia terminated mostly by a sterile cell ( Hollenberg 1969, Tanaka and Chihara 1980, Womersley 1987, Parente and Saunders 2019). Analipus and Heteroralfsia are distinguished by having erect sporophyte and unangium sessile ( Kjellman 1889, Nelson 1980, Kawai 1989, Lim et al. 2007). Fissipedicella is characterized by combinations of these characteristics listed in Table 2 and distinguished from other members ( Analipus, Endoplura, Heteroralfsia, and Ralfsia) within the Ralfsiaceae.
Key to genera within Ralfsiaceae
1. Plants with heteromorphic thalli, crustose portion bearing plurangia, and erect sporophytic thalli bearing unangia ........................................................... Heteroralfsia 1. Plants with isomorphic thalli, either crustose or erect thalli bearing both plurangia and unangia ...................... 2 2. Thalli erect ..................................................... Analipus 2. Thalli crustose ............................................................ 3 3. Chloroplasts per cell several, plurangial reproductive filaments 2–5 ...................................................... Endoplura 3. Chloroplast per cell single, plurangial reproductive filaments 1–2 ........................................................................... 4 4. Tufts of hair in pits present, stalk composed of 1–6 vertically or obliquely cleaved cells ..... Fissipedicella 4. Tufts of hairs in pits absent, stalk composed of 1–2 horizontally cleaved cells ................................ Ralfsia
In the classification of crustose brown algae, the detailed morphology of unangia has been used as a character of various taxonomic ranks within the Ralfsiales ( Tanaka and Chihara 1981b, León-Alvarez and González-González 2003, Lim et al. 2007, León-Alvarez et al. 2017, Parente et al. 2021). Due to the variation in detailed morphologies of unangia, such as stalk-like filaments, paraphyses, and position of unangia, these reproductive structures may hold taxonomic significance within different groups at various levels within the Ralfsiales. There are two main types of unangia: those with paraphyses and those without. Unangia with paraphyses has been observed in the Neoralfsiaceae Lim & Kawai in Lim, Sakaguchi, Hanyuda, Kogame, Phang & Kawai, Pseudoralfsiaceae Patente, Fletcher & G. W. Saunders, Sungminiaceae, and Ralfsiaceae ( Hollenberg 1969, Lim et al. 2007, Parente et al. 2021, Oteng’o et al. 2022). Fissipedicella orientalis produces unangia from stalks with paraphyses, similar to some species of Ralfsia ( Ralfsia confusa Hollenberg, R. hesperia Setchell & N. L. Gardner, R. huanghaiensis Li & Li, R. integra Hollenberg, and R. pedicellata J. Tanaka & Chihara) within Ralfsiaceae ( Hollenberg 1969, Tanaka and Chihara 1981a, Xiyi and Junfeng 1993). However, F. orientalis differs in that it has biseriate stalks formed through vertical or oblique divisions, whereas Ralfsia species have uniseriate stalks formed only through transverse divisions. The development of stalk by the vertical division has not been reported in any other genera within the Ralfsiaceae. Applying molecular analyses to crustose brown algae has significantly broadened our comprehension of the phylogenetic relationships among related families within Ralfsiales, leading to valuable insights into their taxonomic positions. In our molecular analyses, Fissipedicella orientalis is more closely related to incertae sedis (“ Ralfsia longicellularis” and Endopluralean sp. 1GWS) than other genera within Ralfsiaceae. “ Ralfsia longicellularis” was originally described from Peter the Great Bay, Far East Russia by Perestenko (1980) and was subsequently reported to Korea based on molecular and morphological evidence by Oteng’o and Won (2020). “ Ralfsia longicellularis” is clearly distinguished as a different species from other species of Ralfsia in our phylogenetic trees. Endopluraean sp. 1GWS has been reported as a closely related taxon with Endoplura based on molecular data by Parente and Saunders (2019). The sequence divergences for rbcL and COI-5P between Fissipedicella orientalis and the incertae sedis (“ Ralfsia longicellularis” and Endopluralean sp. 1GWS) were 6.7–6.8 and 17.8%, respectively ( Table 1). The sequence divergence values between genera within Ralfsiaceae have been shown to be ≥5.5 and ≥17.5% for rbcL and COI-5P, respectively ( Table 1). In the future, if more information becomes available about the clade of incertae sedis, it may be recognized as a distinct taxonomic group within Ralfsiaceae. Additionally, none of unpublished and non-identified sequences in GenBank included in these analyses have been related to our new genus, Fissipedicella orientalis gen. et sp. nov.
ACKNOWLEDGEMENTS
This study was supported by a research fund from Chosun University (2022) to T. O. Cho.
CONFLICTS OF INTEREST
The authors declare that they have no potential conflicts of interest.
Fig. 1
Phylogenetic tree of Ralfsiales inferred from the concatenated dataset ( rbcL + 5′ region of cytochrome c oxidase subunit I [COI-5P]). The value above branches = maximum likelihood bootstrap (BS) values in % ≥50, Bayesian posterior probabilities (BPP) ≥0.75. Values of <BS 50 or BPP 0.75 are indicated by hyphens (−). Values of BS 100 or BPP 1.00 are indicated by asterisks (*). GenBank accession numbers of each sequence data set are shown in Supplementary Table S1. Species analyzed in this study are marked in bold font. Branch lengths are proportional to substitution rate. 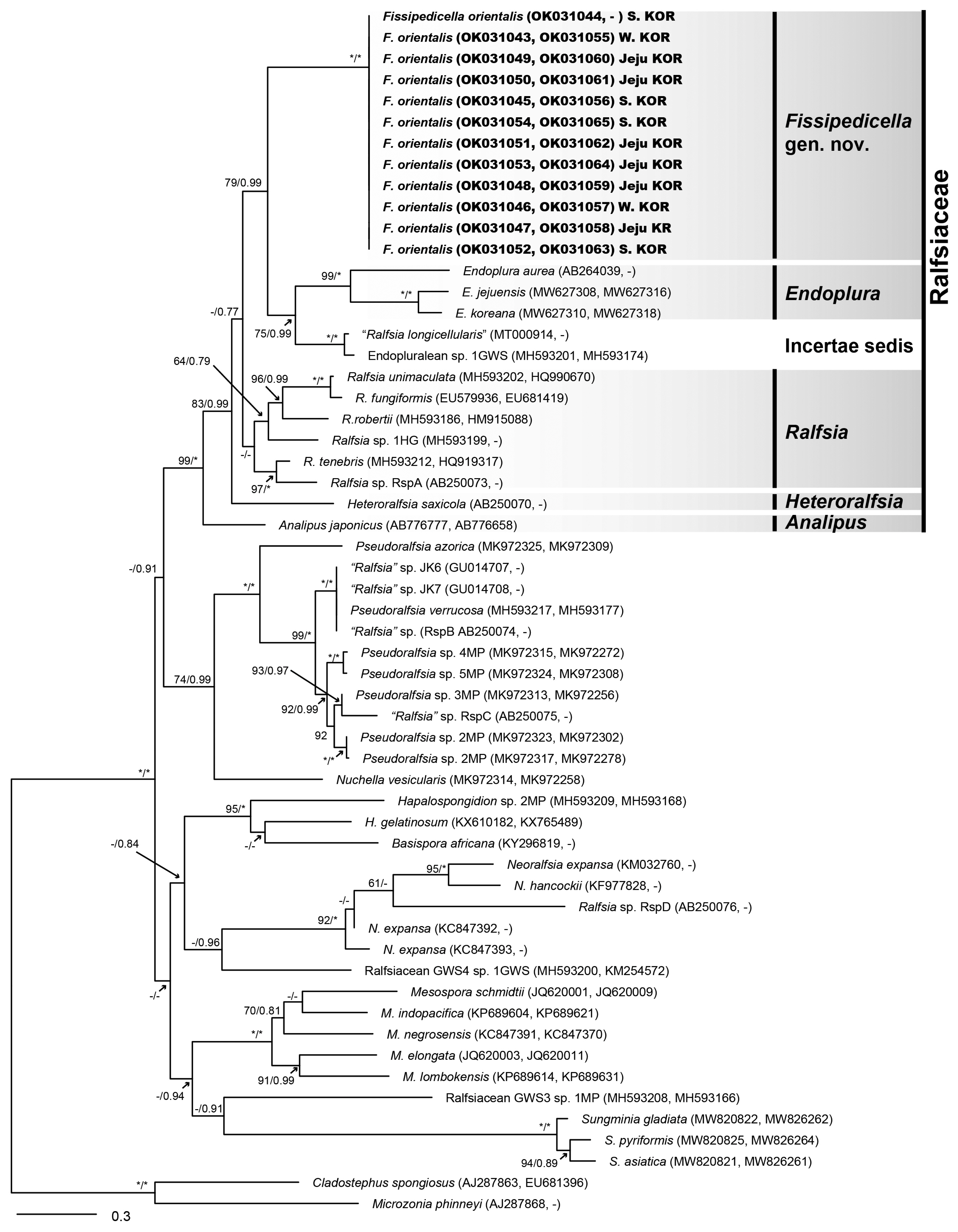
Fig. 2
Morphology of Fissipedicella orientalis sp. nov. (A) Thalli on the rock forming epilithic greenish-brown crusts. (B) Thalli with circular confluent crusts. (C & D) Longitudinal section of vegetative young thallus (C) and mature thallus (D) composed of erect perithallial filaments (EF) and hypothallial basal layer (BL). (E) Erect perithallial filaments showing a single chloroplast (arrows) per cell chloroplasts. (F) Tufts with hairs (Ha) in pits developed from perithallial erect filaments. (G) Longitudinal section view of reproductive thallus showing protrusion of plurangial sorus. (H) Longitudinal section showing reproductive thallus with plurangia (Pl), vegetative filaments (VF), and hypothallial basal layer (BL). (I) Plurangial reproductive structure composed of plurangial filaments (PF) and terminal sterile cells (SC). (J) Longitudinal section of reproductive thallus showing protrusion of unangial sorus. (K) Longitudinal section of reproductive thallus showing unangia (arrow) among paraphyses (Pa). (L) Unangia (Un) on stalks (St) with vertical divisions (arrows) and paraphyses (Pa). Scale bars represent: A, 0.5 cm; B, 0.2 cm; C, D & K, 50 μm; E, 10 μm; F, H, I & L, 25 μm; G & J, 100 μm. 
Table 1
Gene sequence divergence value (%) of rbcL and COI-5P between genera of Ralfsiaceae using pairwise distance
|
Features |
1 |
2 |
3 |
4 |
5 |
6 |
|
1. Fissipedicella gen. nov. |
- |
19.5 |
20.6–21.2 |
- |
17.5–19.3 |
17.8 |
|
2. Analipus
|
7.9 |
- |
19.3–21.4 |
- |
18.0–19.9 |
19.3 |
|
3. Endoplura
|
9.3–10.1 |
9.0–9.5 |
- |
- |
20.4–23.1 |
18.2–19.5 |
|
4. Heteroralfsia
|
9.4 |
7.7 |
10.7–11.1 |
- |
- |
- |
|
5. Ralfsia
|
7.4–8.9 |
5.5–6.7 |
9.3–10.5 |
7.8–9.3 |
- |
16.5–20.1 |
|
6. Incertae sedis |
6.7–6.8 |
7.7–7.9 |
8.1–9.3 |
8.1–8.5 |
6.9–8.1 |
- |
Table 2
Comparison of the morphological characters among genera within Ralfsiaceae
REFERENCES
Abbott, I. A. & Hollenberg, G. J. 1976. Marine algae of California. Stanford University Press, Stanford, CA, 827 pp.
Farlow, W. G. 1881. Marine algae of New England and adjacent coast. US Government Printing Office, Washington, 210 pp.
Guiry, M. D. & Guiry, G. M. 2023. AlgaeBase, World-wide electronic publication, National University of Ireland, Galway, Available from: http://www.algaebase.org
. Accessed Jun 20, 2023 Hollenberg, G. J. 1969. An account of the Ralfsiaceae (Phaeophyta) of California. J. Phycol. 5:290–301.   Kawai, H. 1989. Life history and systematic position of Heteroralfsia saxicola gen. et comb. nov. (Ralfsiaceae, Phaeophyceae). Phycologia. 28:243–251.  Keum, Y. S. 2010. Sphacelariales, Cutleriales, Ralfsiales. In : Shin S., editor Algal Flora of Korea. Vol. 2, No 2 Heterokontophyta: Phaeophyceae: Ishigeales, Dictyotales, Desmarestiales, Sphacelariales, Cutleriales, Ralfisales, Laminariales. National Institute of Biological Resources, Incheon, 73–108.
Kjellman, F. R. 1889. Om Beringhafvets algflora [About the Bering Sea algae]. Kongl. Sv. Vet. Acad. Handl. Stockholm. 23:1–58.
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. 2016. PartitionFinder 2: new methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34:772–773.   Lee, I. K. & Kang, J. W. 1986. A check list of marine algae in Korea. Korean J. Phycol. 1:311–325.
Lee, Y. 2008. Marine algae of Jeju. Academy Publication, Seoul, 477 pp.
Lee, Y. P. & Kang, S. Y. 2001. A catalogue of the seaweeds in Korea. Jeju National University Press, Jeju, 662 pp.
León-Alvarez, D. & González-González, J. 2003. The morphological distinction of Ralfsia expansa and R. hancockii (Ralfsiaceae, Rhaeophyta) from Mexico. Phycologia. 42:613–621.  León-Alvarez, D., Reyes-Gómez, V. P., Wynne, M. J., Ponce-Márquez, M. E. & Quiróz-González, N. 2017. Morphological and molecular characterization of Hapalospongidion gelatinosum, Hapalospongidiaceae fam. nov. (Ralfsiales, Phaeophyceae) from Mexico. Bot. Mar. 60:567–581.
Lim, P.-E., Sakaguchi, M., Hanyuda, T., Kogame, K., Phang, S.-M. & Kawai, H. 2007. Molecular phylogeny of crustose brown algae (Ralfsiales, Phaeophyceae) inferred from rbcL sequences resulting in the proposal for Neoralfsiaceae fam. nov. Phycologia. 46:456–466.  Nelson, W. A. 1980.
Analipus japonicus (Harv.) Wynne (Phaeophyta): studies of its biology and taxonomy. Ph.D. dissertation. University of British Columbia, Vancouver, Canada, 244 pp.
Oteng’o, A. O., Cho, T. O. & Won, B. Y. 2021.
Endoplura jejuensis sp. nov. and E. koreana sp. nov. (Ralfsiales, Phaeophyceae) from Korea based on molecular and morphological analyses. Algae. 36:155–163.   Oteng’o, A. O. & Won, B. Y. 2020.
Ralfsia longicellularis (Ralfsiales, Phaeophyceae): a Far East Asian endemic brown alga from Korea. Korean J. Environ. Biol. 38:101–105.  Parente, M. I., Fletcher, R. L., Costa, F. O. & Saunders, G. W. 2021. Taxonomic investigation of Ralfsia-like (Ralfsiales, Phaeophyceae) taxa in the North Atlantic Ocean based on molecular and morphological data, with descriptions of Pseudoralfsiaceae fam. nov., Pseudoralfsia azorica gen. et sp. nov. and Nuchella vesicularis gen. et sp. nov. Eur. J. Phycol. 56:12–23.  Parente, M. I. & Saunders, G. W. 2019. A molecular survey of Ralfsia sensu stricto (Ralfsiales, Phaeophyceae) in Canada uncovers three new species: R. robertii sp. nov., R. tenebris sp. nov., and R. unimaculata sp. nov. Botany. 97:135–147.
Perestenko, L. 1980. Vodorosli Zaliva Petra Velikogo [The seaweeds of Peter the Great Bay]. NAUKA Leningradskoe Otdelenie, Leningrad, 231 pp.
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. 2018. Posterior summarization in Bayesian phylogenetic using Tracer 1.7. Syst. Biol. 67:901–904.   Setchell, W. A. & Gardner, N. L. 1924. Phycological contributions, VII. Univ. Calif. Publ. Bot, 13:1–13.
Silberfeld, T., Rousseau, F. & de Reviers, B. 2014. An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptogam. Algol. 35:117–156.  Silvestro, D. & Michalak, I. 2012. raxmlGUI: a graphical front-end for RaxML. Org. Divers. Evol. 12:335–337.   Tanaka, J. & Chihara, M. 1980. Taxonomic study of the Japanese crustose brown algae (2). Ralfsia (Ralfsiaceae, Ralfsiales) (Part 1). J. Jpn. Bot. 55:337–342.
Tanaka, J. & Chihara, M. 1981a. Taxonomic study of the Japanese crustose brown algae (4) Ralfsia (Ralfsiaceae, Ralfsiales) (Part 3). J. Jpn. Bot. 56:97–104.
Tanaka, J. & Chihara, M. 1981b. Taxonomic study of Japanese crustose brown algae (5). Endoplura and Diplura (Ralfsiaceae, Ralfsiales). J. Jpn. Bot. 56:153–160.
Womersley, H. B. S. 1987. The marine benthic flora of southern Australia Part II. South Australian Government Printing Division, Adelaide, 484 pp.
Xiyi, L. & Junfeng, L. 1993.
Ralfsia huanghaiensis, a new species of Ralfsia (Ralfsiales, Phaeophyta). Ocenol. Limnol. Sin. 24:172–176.
|
| TOOLS |
 |
 PDF Links PDF Links |
 |
 PubReader PubReader |
 |
 ePub Link ePub Link |
 |
 Full text via DOI Full text via DOI |
 |
 Download Citation Download Citation
|
 |
 Supplement1 Supplement1
 Supplement2 Supplement2
|
 |
 Print Print
|
 |
Share: 



|
 |
METRICS  |
|
|
|
| Related articles |
| Endoplura jejuensis sp. nov. and Endoplura koreana sp. nov. (Ralfsiales, Phaeophyceae) from Korea based on molecular and morphological analyses 2021 September;36(3) |
| Platysiphon verticillatus and Arcticophycus glacialis: gen. et comb. nov. (Stschapoviales, Phaeophyceae) based on ecological considerations, life history, morphology, and ultrastructure 2019 September;34(3) |
| Alsidium oliveiranum sp. nov. (Rhodomelaceae, Rhodophyta), an overlooked species from the southwestern Atlantic based on morphology and DNA sequence data 2019 September;34(3) |
| Viator vitreocola gen. et sp. nov. (Stylonematophyceae), a new red alga on drift glass debris in Oregon and Washington, USA 2019 June;34(2) |
| A novel species Symphyocladia glabra sp. nov. (Rhodomelaceae, Rhodophyta) from Korea based on morphological and molecular analyses
2013 ;28(2) |
|
|













