INTRODUCTION
Establishing a practical taxonomic system is the primary method to identify and understand organismal diversity. However, this system has been constantly adjusted along the introduction of new methods and new progress of modern science. Cyanobacteria are one of the most important phyla of photosynthetic prokaryotes with large morphological and metabolic diversity, and they are widely distributed on the earth, including extreme environments (Castenholz and Waterbury 1989). The taxonomy of this phylum had been largely based on morphological characteristics which were often insufficient to define genera and species (Komárek 2003, Taton et al. 2003, 2006, Johansen and Casamatta 2005, Turicchia et al. 2009, Genuário et al. 2013). Moreover, some morphological characters vary under different conditions, such as colonial formation, cellular size, presence of sheaths, heterocytes and akinetes (Hašler et al. 2012, Dvořák et al. 2015a, Hentschke and Sant’Anna 2015, Caires et al. 2018). Berrendero et al. (2011) found that morphological features of Calothrix changed in the culture condition, making them to look like Tolypothrix. To avoid these issues arising from morphological plasticity, phylogenetic relationships among cyanobacteria at all catagories have been inferred using the 16S rRNA gene (Giovannoni et al. 1988, Turner et al. 2001, Suda et al. 2002, Komárek and Kaštovský 2003) since it is highly conserved among all prokaryotes (Coenye and Vandamme 2003). Further, the polyphasic approach, based on morphological, physiological and ecological and molecular characters, has been proposed to better improve cyanobacterial taxonomic assignments and revisions, therefore resulting in the description of numerous cyanobacterial genera and species (Fiore et al. 2007, Bohunická et al. 2015, Genuário et al. 2015, Sciuto and Moro 2016, Sciuto et al. 2017).
The family Nostocaceae traditionally consisted of the heterocytous taxa without false branching or tapering filaments (Geitler 1932, Komárek 2013), and Nostocaceae was characterized by morphological features, such as isopolar filaments and usual formation of akinetes. However, numerous morphologically well-defined genera in this family, such as Anabaena, Trichormus, and Wollea, were shown to be polyphyletic based on molecular phylogenetic analyses (Rajaniemi et al. 2005a, Kozhevnikov and Kozhevnikova 2011, Zapomělová et al. 2013). Solving such problems through carefully taxonomic re-examination and phylogenetic clustering could lead to the description of new genera / species.
Nostoc, the type genus of this family, is the largest but most clearly polyphyletic genus, and many previous studies already indicated the wide genetic diversity of this genus (Hrouzek et al. 2005, Řeháková et al. 2007, Kaštovský and Johansen 2008, Lukešová et al. 2009). The formation of mucilaginous colonies and the complex life cycle are major traits common to all species in the Nostoc genus. In spite of its well-established morphological features, Nostoc has been proven to be genetically heterogeneous, and several genotypic groups were shown to fall outside of the ‘Nostoc sensu stricto’ clade based on 16S rRNA gene phylogeny (Hrouzek et al. 2005, 2013, Rajaniemi et al. 2005a, Papaefthimiou et al. 2008, Lukešová et al. 2009, Silva et al. 2014). Therefore, the genus Nostoc has been proposed to be amended in consideration of phylogenetic data, and some Nostoc-related morphotypes have been established as new genera including Mojavia, Desmonostoc, Halotia, Komarekiella, Aliinostoc, Compactonostoc, and Minunostoc (Řeháková et al. 2007, Hrouzek et al. 2013, Genuário et al. 2015, Bagchi et al. 2017, Hentschke et al. 2017, Cai et al. 2019a, 2019b).
During a broader survey on terrestrial heterocytous cyanobacteria in China, we isolated two strains of morphologically distinct heterocytous cyanobacteria from a seldom sampled subtropical region, the Small Seven-hole Scenic Area, in Guizhou province. In this study, we characterized these two strains using a polyphasic approach combining morphological, ecological, and molecular analyses. These two strains bear morphological characteristics resembling Nostoc in successive stages of their life cycle. However, they did not fall within the main Nostoc cluster based on a phylogenetic analysis. The purpose of this paper is to establish the phylogenetic position of these two strains, distinguish them from other cyanobacterial genera and demonstrate that they belong to a novel genus, Violetonostoc, and the type species Violetonostoc minutum sp. nov. is here described.
MATERIALS AND METHODS
Sampling and cultivation
Environmental samples were collected from Small Seven-hole Scenic Area (25°15′34.79″ N, 107°42′46.72″ E), in Guizhou province, China in October 2014. This is a place with subtropical humid monsoon climate, and the air temperature was 12°C. A visible growth of cyanobacteria on rocky substrates was collected with a ladle. Using the micropipette washing method (Rippka 1988) to isolate the heterocytous filaments into unialgal cultures, two strains of Nostoc-like cyanobacteria were obtained and cultured in screw-capped glass tubes containing 6 mL of BG11 medium. The strains were kept at 25°C under a 12 h : 12 h LD cycle with a photon flux density of 35 μmol m−2 s−1 from white fluorescent lamps. The living culture were maintained in the Chinese Harmful Algae Biology (CHAB) culture collection of the Institute of Hydrobiology, China. The serial number of the studied strains are CHAB 5840 and CHAB 5841. The dry material of strain CHAB 5840 was obtained by freeze-drying at −40°C and stored at the Freshwater Algal Herbarium (HBI), Institute of Hydrobiology, Chinese Academy of Science, Wuhan, China.
Morphological characterization
Morphometric characters, including length and width of vegetative cells and heterocytes were measured from ≥50 individuals using a Nikon Eclipse 80i light microscope with DS-Ri1 digital camera (Nikon, Tokyo, Japan). The images were analyzed using the NIS-Elements D 3.2 with the magnification of microscopy as 600× and 1,000×, respectively, and the microscopic observation was conducted in the brightfield.
DNA extraction and polymerase chain reaction amplification
Unialgal cultures of these two strains were harvested and the total genomic DNA was extracted using the modified cetyltrimethylammonium bromide method employed by Neilan et al. (1995). The primers PA and B23S (Edwards et al. 1989, Gkelis et al. 2005) were used for amplification of 16S rRNA gene, and the primers 322 and 340 (Iteman et al. 2000) were used to amplify the 16S–23S internal transcribed spacer (ITS) rRNA. Polymerase chain reaction (PCR) was performed in an MJ Mini Personal Thermal Cycler (Bio-Rad, Hercules, CA, USA), and the amplification program was carried out as follows: one cycle of 3 min at 94°C; 34 cycles of 30 s at 94°C, 30 s at 58°C (30 s at 55°C for ITS), and 1 min at 72°C (30 s for ITS) and then a final 5 min elongation step at 72°C. The PCR amplification products were purified by QIAquik PCR purification columns (Qiagen, Hilden, Germany) according to manufacturer’s manual and then cloned into the pMDTM18-T vector (TaKaRa, TaKaRa Bio Inc., Otsu, Japan). The cloning procedure was performed following by Sambrook and Russell (2001). Sanger dideoxy sequencing was carried out by using an ABI 3730 Automated Sequencer (PerkinElmer, Waltham, MA, USA).
Phylogenetic analysis
Sequences for phylogenetic analyses were downloaded from GenBank based on a BLAST search. 16S rRNA gene sequences were aligned using CLUSTAL X ver. 2.0 (Larkin et al. 2007). The final phylogenetic trees were constructed using neighbor-joining (NJ), maximum likelihood (ML), and bayesian inference (BI). The NJ analysis using Kimura-2 model upon default parameters with 1,000 bootstrap replicates were run via MEGA software v7.0 (Kumar et al. 2016), the ML algorithms were performed using PhyML V3.0 (Guindon et al. 2010) and BI were analyzed with MrBayes 3.2.2 (Ronquist et al. 2012). The ModelFinder (Kalyaanamoorthy et al. 2017) program was used to explore the sequence evolution model that fitted the dataset based on Akaike information criterion. The best-fit models, selected for the ML and BI analyses of the 16S rRNA gene were GTR + I + G, the obtained phylogenetic tree was edited with FigTree v1.4.3 (Rambaut 2016). Gloeobacter violaceus VP3-01 was chosen as the outgroup taxon for 16S rRNA phylogeny. Calculation of p-distance was done with MEGA software v7.0 (Kumar et al. 2016) and used to calculate sequence identity [100 × (1 - p)] for 16S rRNA data.
Accession numbers for GenBank sequences that were used for phylogenetic analyses were listed as a supplementary file (Supplementary Table S1). The obtained nucleotide sequences of our sequences have been deposited in the GenBank database with the accession numbers: MN400069, MN400070, MN400067, and MN400068. 16S–23S rRNA secondary structures of D1-D1′, Box-B, and V3 helices were determined using “RNAstructure”, ver. 5.6 (Mathews Lab 2013).
RESULTS
Violetonostoc F. Cai & R. Li gen. nov
Description
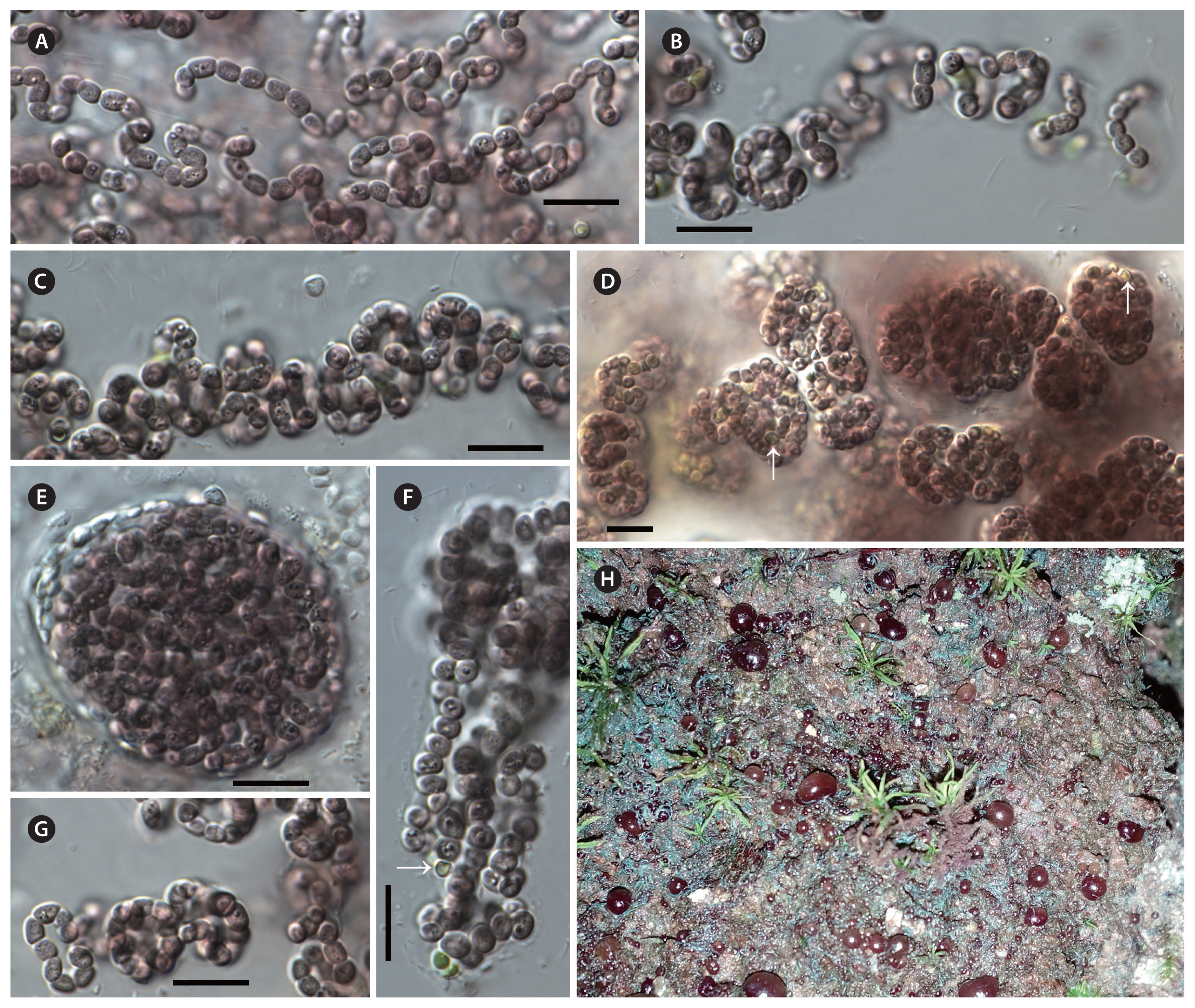
Thallus macroscopic, free-living on rocky substrates in natural conditions, violet to black-brown. In liquid culture, growing as a spherical macroscopic biomass on the bottom of the test tube. Long filaments loosely aggregated in young colonies, and always enclosed by firm mucilage layer. The young colonies eventually becoming compact macroscopic colonies with densely entangled filaments. Sheath colorless, usually visible in younger colonies. Trichomes isopolar, apical cell morphologically not different from other cells. Cells spherical to barrel-shaped. Heterocytes, spherical or oval, smaller than vegetative cells, terminal in young colonies, intercalary in old colonies. Akinetes not known. Reproduction by fragmentation of the trichomes into motile hormogonia.
Violetonostoc minutum F. Cai & R. Li sp. nov. (Fig. 1)
Description
In culture, growing as a spherical macroscopic biomass on the bottom of the test tube or attached to the tube walls. Long filaments loosely aggregated in young colonies, and always enclosed by firm or wide mucilage layer. The young colonies eventually becoming compact spherical macroscopic colonies, with densely entangled filaments. Sheath colorless, usually distinct at younger colony margins, and not visible in the older colonies. Cells violet to black-brown, sometimes granulated, spherical to barrel-shaped, 2.17–2.81 μm long and 1.8–2.6 μm wide (smaller than that of Nostoc-like genera). Heterocytes spherical or oval, smaller than vegetative cells, 1.15–1.85 μm long, 0.95–1.76 μm wide (smaller than that of Nostoc-like genera), terminal in young colonies, intercalary in old colonies. Akinetes not observed during three years of cultivation.
Holotype
Dry material of the strain CHAB 5840 was stored at the Freshwater Algal Herbarium (HBI), Institute of Hydrobiology, Chinese Academy of Science, Wuhan, China, as specimen No. SSSA201402.
Type locality
Small Seven-hole Scenic Area (25°15′34. 79″ N, 107°42′46.72″ E), Guizhou province, China.
Molecular and phylogenetic analysis
The evolutionary distance based on the 16S rRNA gene showed that two Violetonostoc strains shared 99.6% similarity with each other (Table 1); they shared 94.3–95.2% similarity with Nostoc, 93.4–93.5% similarity with Aliinostoc, 95.3–95.7% similarity with Desmonostoc, 94.6% similarity with Halotia, Mojavia, and Minunostoc, 96.1% similarity with Komarekiella and Compactonostoc.
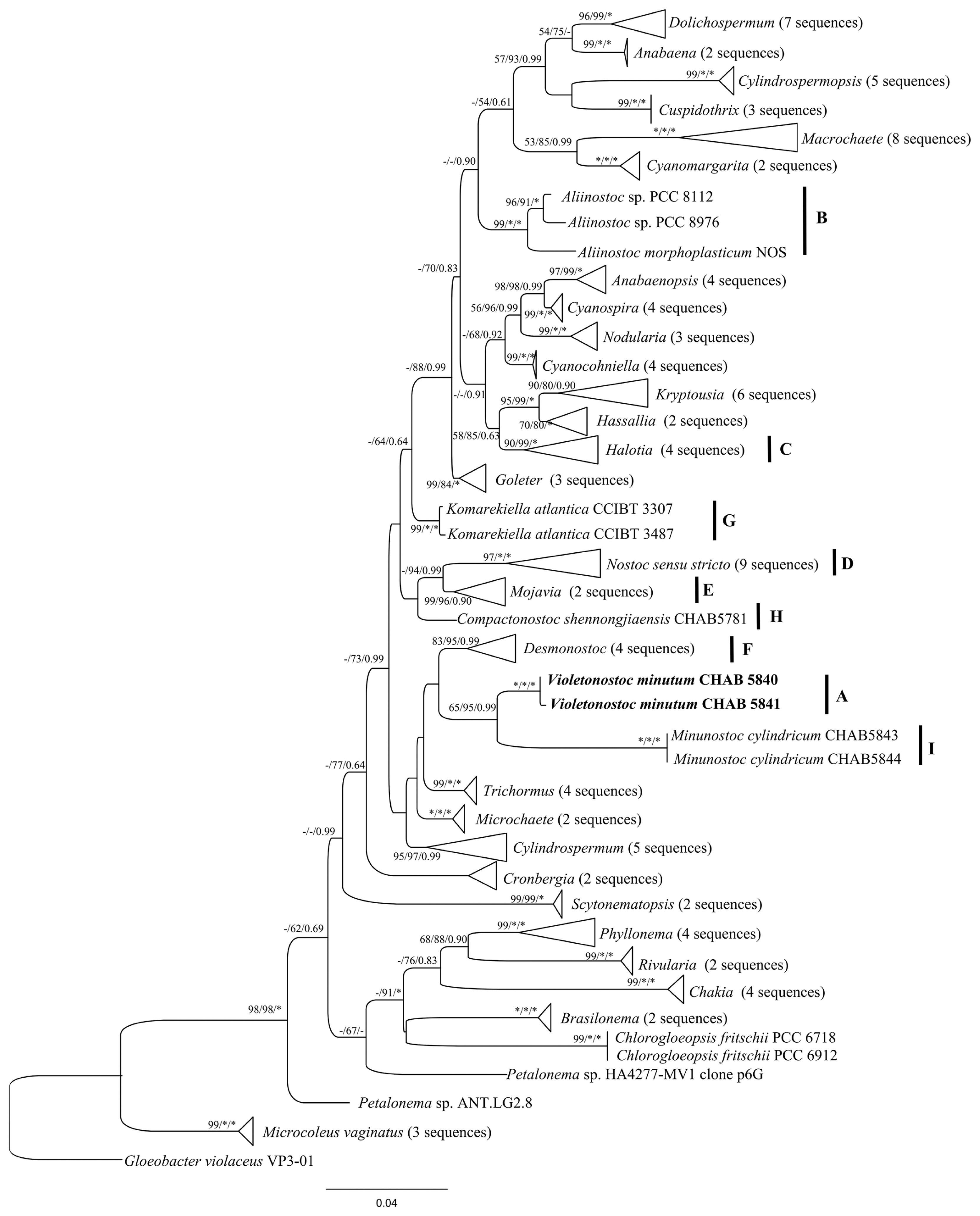
The phylogenetic trees of NJ, ML, and BI based on 16S rRNA gene sequences from 118 cyanobacterial taxa including CHAB 5840 and CHAB 5841 were constructed. The phylogenetic analysis (ML tree) (Fig. 2) exhibited that two strains of Violetonostoc were grouped into a unique clade, which was supported by NJ, ML, and BI approaches with bootstrap values of 100%, 100%, and 1.00, respectively. And these two strains were distinct from the Nostoc sensu stricto clade, a phylogenetically close taxon was composed of Desmonostoc and Minunostoc group strains. Phylogenetic trees presented good support for genera, especially for Nostoc sensu stricto clade, in which the strains CHAB 5840 and CHAB 5841 are included, as well as Komarekiella, Mojavia, Aliinostoc, Halotia, Desmonostoc, Compactonostoc, and Minunostoc.
ITS secondary structures
In total, three Nostoc species, two Desmonostoc species, one Halotia, Mojavia, Komarekiella, Goleter, Trichormus, Aliinostoc, Compactonostoc, Minunostoc species and Violetonostoc (CHAB 5840, CHAB 5841) were used to compare the secondary structures within Nostocaceae.
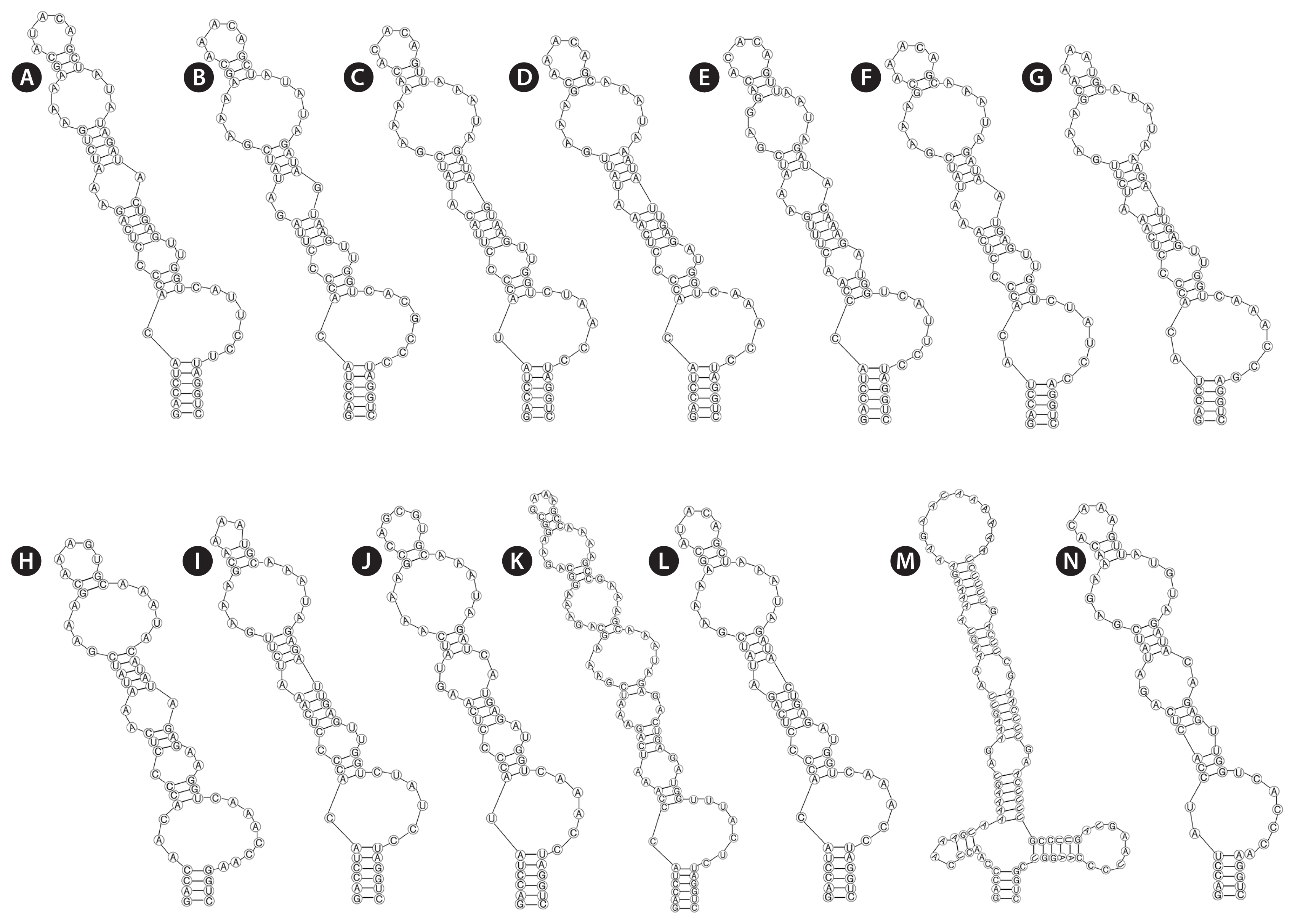
Analyses of the secondary structures of D1-D1′ helix (Fig. 3) showed similar patterns in several genera, except for Minunostoc, which had oddly branched side loop with two branches at the base of the stem, and other closely genera do not possess this unique structure (Fig. 3M). D1-D1′ helix of the two strains consisted of a 5 bp helix (6 bp in Nostoc, Desmonostoc, Aliinostoc, and Compactonostoc; 4 bp in Komarekiella and Minunostoc), followed by a 2 : 7 bp base bilateral bulge (1 : 6 [7] base bilateral bulges in Nostoc, Desmonostoc, Aliinostoc, and Compactonostoc; 2 : 6 and 3 : 8 base bilateral bulges in Mojavia and Komarekiella, respectively), and then by a unpaired nucleotide (U) on 3′ side of their basal unilateral bulges (other genera do not possess unpaired nucleotides); following this side loop were three bilateral bulges (four bilateral bulges in Minunostoc and Komarekiella, six bilateral bulges in Aliinostoc).
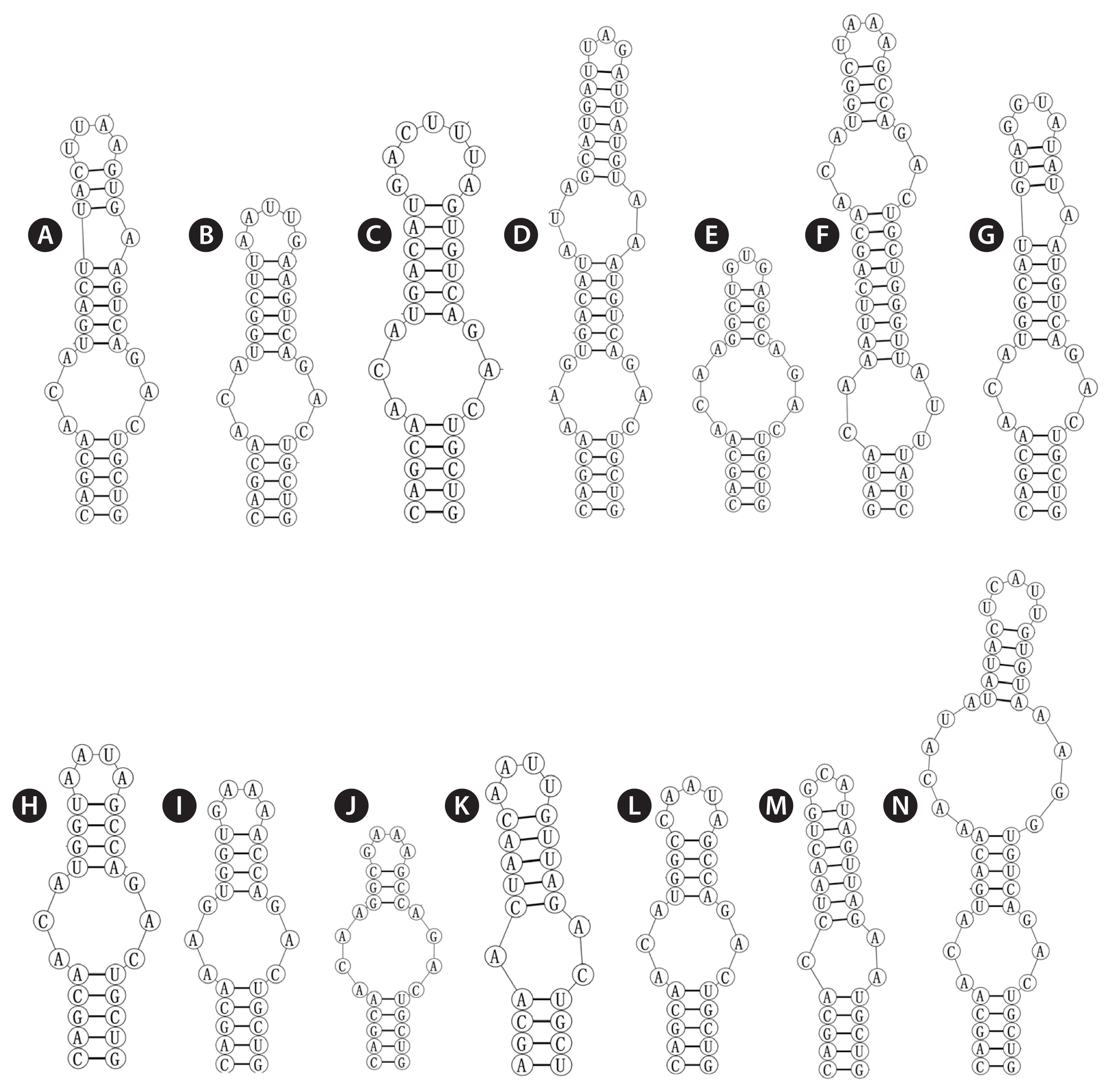
The base stem of the Box-B helix of Violetonostoc (Fig. 4N) consisted of 5 bp helix, followed by a 3 : 3 base bilateral bulge, and then further followed by a large 6 : 4 base bilateral bulge, the terminal loop contained 5 bp bases (UCAUU), which was differed from other taxa.
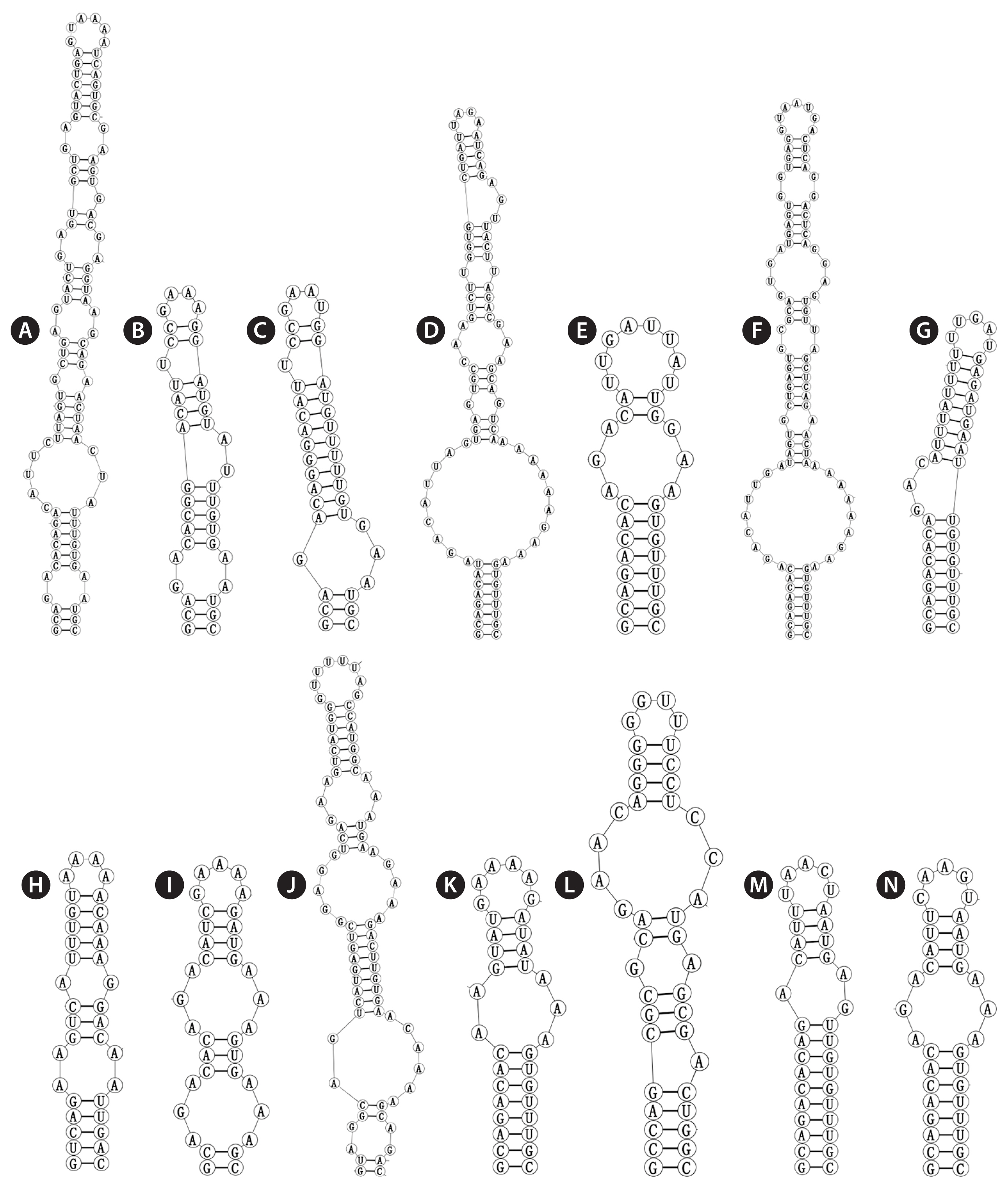
The V3 of Violetonostoc was distinct from the V3 helix in all other taxa in the analysis (Fig. 5N), the base of the stem of CHAB 5840 and CHAB 5841 consisted of an 8 bp helix, followed by a 3 : 3 base bilateral bulge, and then further followed by the terminal loop consisting of a 5 bp nucleosides (CAAGU). The basal stem of V3 in Nostoc species contains a 3 bp helix and then followed by one or two bilateral bulges (six in strains Nostoc lichenoides CNP-AK1), three types were presented in three Nostoc species (Fig. 5A & C), and two types were observed in two Desmonostoc species (Fig. 5D & E). The V3 helix of recently established genera were also quite different within each other, the base of the stem of Mojavia pulchra JT2-VF2 consisted of 8 bp helix, then further followed by five bilateral bulges (Fig. 5F), the basal stem of V3 in Halotia branconii CENA392 contained a 9 bp helix, then further followed by a side loop with three unpaired bases on the 5′ side (Fig. 5G). While Komarekiella atlantica CCIBT 3481 contained a 5 bp helix, and then followed by two bilateral bulges (Fig. 5H), and the base of the stem of Aliinostoc morphoplasticum NOS consisted of 8 bp helix, followed by bilateral bulge of 2 : 3 bases, then further followed by the terminal loop consisting of a 6 bp nucleosides (CAAAAG) (Fig. 5K). The base stem of Compactonostoc consisted of a 5 bp helix (GCCAG), followed by a side loop with a single base on the 3′ side, and then by one 4 : 3 bases bilateral bulge and the terminal loop consisting of a four nucleotide bases (GGUU) (Fig. 5L). V3 helix of Minunostoc consisted of very long stems (10 bp), followed by a 1 : 2 base bilateral bulge, and further followed by a 5 bp terminal loop (Fig. 5M).
DISCUSSION
Previous studies repeatedly indicated that using few morphological characteristics for cyanobacterial taxonomy have been proven to underestimate its biodiversity (Sherwood et al. 2014, Dvořák et al. 2015a, Shalygin et al. 2017). Many cyanobacteria from diverse categories (at specific, generic, and familial level) have been found to be polyphyletic (Wilmotte and Golubić 1991, Wilmotte and Herdman 2001, Gugger et al. 2002a, 2002b, Hoffmann et al. 2005, Rajaniemi et al. 2005a, 2005b, Komárek et al. 2014). It is generally accepted that cyanobacterial genera must be monophyletic clusters in the newest cyanobacterial taxonomy (Komárek et al. 2014, Komárek 2018).
The nostocacean cyanobacteria have more or less complicated filaments and also largely encountered a considerable revision at taxonomic categories. Nostocaceae is among one of the most problematic group, and several genera in this family were arguably polyphyletic (Rajaniemi et al. 2005a, Kozhevnikov and Kozhevnikova 2011, Zapomělová et al. 2013, Komárek et al. 2014, Choi et al. 2018). Moreover, a number of strains assigned to Tolypothrix, Calothrix, and Microchaete have been found to form a clade within this family (Kaštovský and Johansen 2008, Kaštovský et al. 2014, Komárek et al. 2014). The genus Nostoc was morphologically defined as having unbranched uniseriate isopolar filaments with differentiated akinetes and terminal / intercalary heterocytes embedded in mucilaginous material-forming colonies. It is becoming difficult to clearly differentiate closely related taxa based solely on morphology owing to the huge amount of heterogeneity (Singh et al. 2016). Furthermore, molecular data indicate that the genetic diversity within this genus exceeds its morphological diversity, clearly supporting its polyphyletic status (Rajaniemi et al. 2005a, 2005b, Řeháková et al. 2007, Hrouzek et al. 2013, Genuário et al. 2015, Bagchi et al. 2017, Hentschke et al. 2017). Therefore, the genus Nostoc requires continuously taxonomic re-evaluation and the establishment of diacritical features. The traditional definition of Nostoc includes several distinct but not closely related phylogenetic clusters, some of which have been already formally separated from Nostoc, such as Mojavia (Řeháková et al. 2007) and Desmonostoc (Hrouzek et al. 2013). In recent years, many studies using the polyphasic approach led to the establishment of more novel genera by splitting from Nostoc (Genuário et al. 2015, Bagchi et al. 2017, Hentschke et al. 2017, Cai et al. 2019a, 2019b). In China, the investigation on cyanobacteria from all kinds of environmental conditions and researches on cyanobacterial taxonomy have been largely performed (Song et al. 2015a, 2015b, Li and Li 2016, Li et al. 2017, Cai et al. 2018, 2019a, 2019b, Wang et al. 2019). The present study described Violetonostoc as a new cyanobacterial genus isolated from the Small Seven-hole Scenic Area in Guizhou province, and this is the second new Nostoc-like genus from the same site, following the first new one as Minunostoc (Cai et al. 2019a). The two strains in this study are morphologically similar, and it is not possible to separate them as different species. The molecular data also showed very high similarities in the 16S rRNA gene sequence (99.6%) and the 16S–23S ITS gene region (100%) between both strains. Morphologically, this novel genus initially seemed to belong to Nostoc, whose filaments are unbranched and always embedded in mucilage. However, the molecular and phylogenetic results confirmed the polyphyletic status of the Nostoc-like cyanobacteria and supported its separation from the Nostoc sensu stricto clade, allowing us to describe these two strains as a new genus.
Based on the phylogeny of 16S rRNA gene (Fig. 2), these two Nostoc-like strains were grouped into a unique and compacted cluster with high bootstrap values to distantly separate from clade D consisted of nine Nostoc strains including the type species Nostoc commune. Clade I consisting of two strains of Minunostoc cylindricum was the sister taxon. As shown in Fig. 2, the phylogenetic tree based on 16S rRNA gene sequences revealed that several nostocacean genera resulted in the formation of 13 clusters, and the Nostoc-like morphotype were phylogenetically divided into clades A, B, C, E, F, G, H, and I, corresponding to genera Violetonostoc, Aliinostoc, Halotia, Mojavia, Desmonostoc, Komarekiella, Compactonostoc, and Minunostoc, respectively.
Furthermore, on the basis of the evolutionary distance matrix, the 16S rRNA gene sequences of Violetonostoc had over 95% similarities to representatives of the following well-established genera: Komarekiella 96.1%, Compactonostoc 96.1%, Trichormus 95.5%, Desmonostoc 95.3–95.7%. While they had less than 95% similarities to the phylogenetically related genus Minunostoc. In bacteriology, using percentage identities of less than 95% 16S rRNA gene sequences to establish genera has been recommended by some researchers (Wayne et al. 1987, Stackebrandt and Goebel 1994), but this sharp limit cannot be used as an absolute criterion for separation of genera for most cyanobacteria. Some studies have recognized the proposal of new genera with higher percentage of 16S rRNA sequence similarities to the existing genera as long as phenotypic or phylogenetic evidence exists for such recognition. Previous studies already demonstrated that genera in Nostocaceae shared higher similarities in the 16S rRNA gene sequence, as frequently more similar than 96% (Flechtner et al. 2002, Řeháková et al. 2007, Berrendero et al. 2011, Kaštovský et al. 2014, Hentschke et al. 2017, Cai et al. 2019b). The phylogenetic analysis revealed that the two strains (CHAB 5840, CHAB 5841) in this study formed a single and unique clade, thus the proposal for the new genus will contribute to solve taxonomic confusion and achieve monophyly within the genus Nostoc.
Setting the new genus Violetonostoc is also supported by the 16S–23S ITS secondary structure. At first glance, the D1-D1′ helix of several genera were quite similar, except for Minunostoc with two unidirectional bulges at the base of the stem and a pronounced, terminal loop, while other closely genera do not possess this unique structure (Cai et al. 2019a). A detailed analysis revealed that the D1-D1′ helix in genus Violetonostoc exposed one unpaired nucleotide (U) on 3′ side of their basal unilateral bulges in comparison to no unpaired nucleotides or one unpaired nucleotide on 5′ side for other taxa. Besides, several unique compensatory base changes are present in the central region of the helix of Violetonostoc. According to these minor variations, this structure is able to separate Violetonostoc from the other genera mentioned above. Moreover, the Box-B and V3 helix showed wide differences in comparison with those most related morphotypes. Consequently, D1-D1′, Box-B, and V3 secondary structures sufficiently separated Violetonostoc from the other genera and thus supported the description of this new genus. Two strains CHAB 5840 and CHAB 5841 shared the same D1-D1′, Box-B and V3 secondary structures.
Morphologically, the species of the genera Violetonostoc and Mojavia are very similar to the species of the true genus Nostoc, with typic morphological features as filaments aggregated and entangled irregularly in gelatinous colonies, and usually shown as macroscopic with peripheral periderm enveloping the whole spherical colonies. A summary of morphological comparisons among Violetonostoc and related genera (Table 2) indicates the difficulty by using morphology alone to define them. The differences for distinguishing Nostoc from Violetonostoc are the cell size and color of thallus. The heterocytes of Nostoc are usually wider than 2 μm, while those of Violetonostoc are usually narrower than 2 μm, and the vegetative cells of Violetonostoc are smaller than those of Nostoc. The monophyly of the Violetonostoc cluster and its phylogenetic separation from the well-defined genera Mojavia, Komarekiella, Compactonostoc, Minunostoc, Nostoc, and Desmonostoc indicate that strains of the genus Violetonostoc share a common ancestor and belong to a novel genus.
This study helps to better understand the diversity of Nostoc-like cyanobacteria and shows higher biodiversity of heterocytous cyanobacteria on some seldom sampled subtropical environments in China which was poorly explored. Cyanobacterial communities in the Small Seven-hole Scenic Area of China remain largely understudied, thus it is necessary to increase sampling in this region, as well as using the polyphasic approach to elucidate cyanobacterial diversity in the future studies.
SUPPLEMENTARY MATERIALS
Supplementary Table S1
List of sequences from the obtained study and NCBI database used in this analysis, with accession numbers (https://e-algae.org).